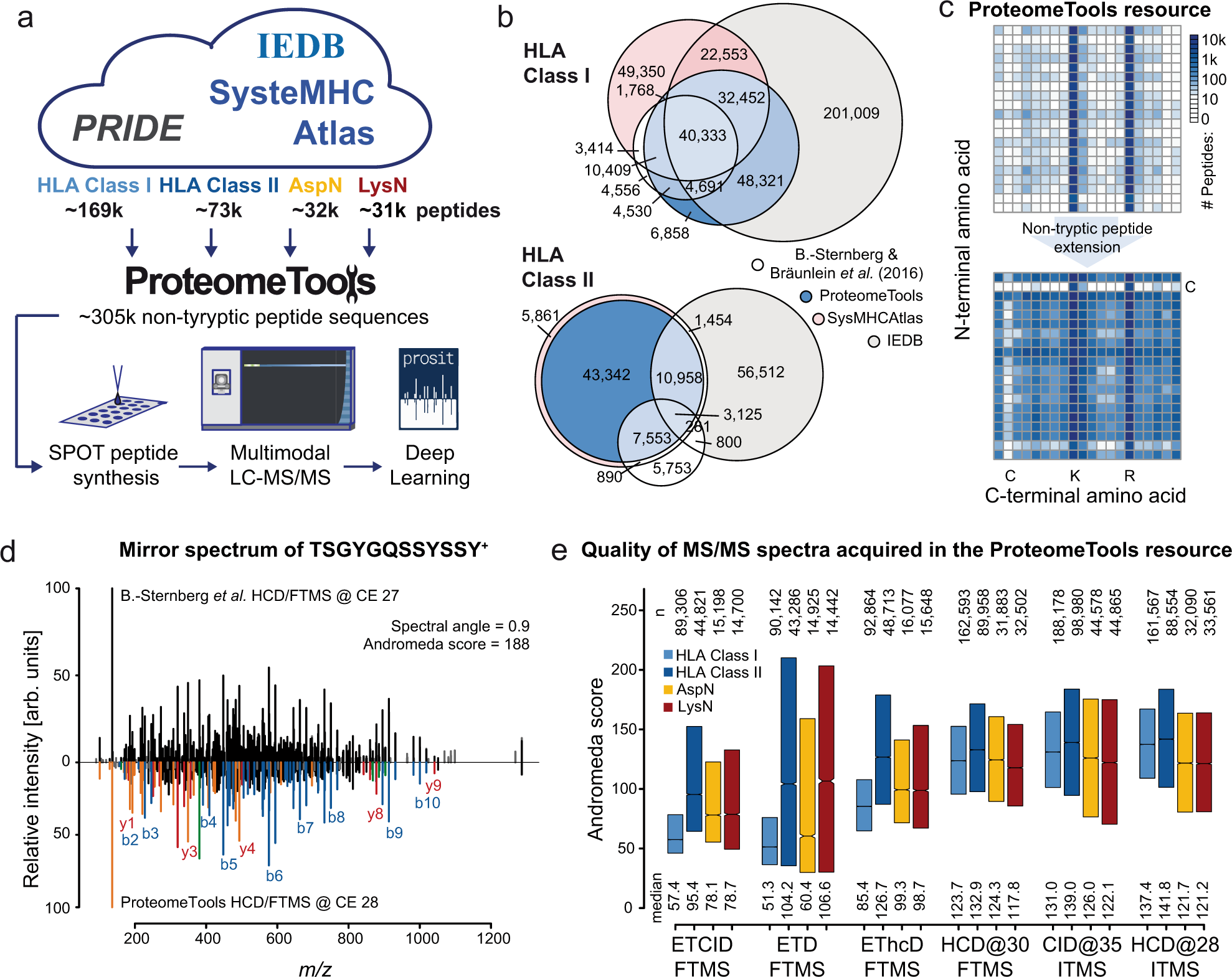
Deep learning boosts sensitivity of mass spectrometry-based immunopeptidomics | Nature Communications
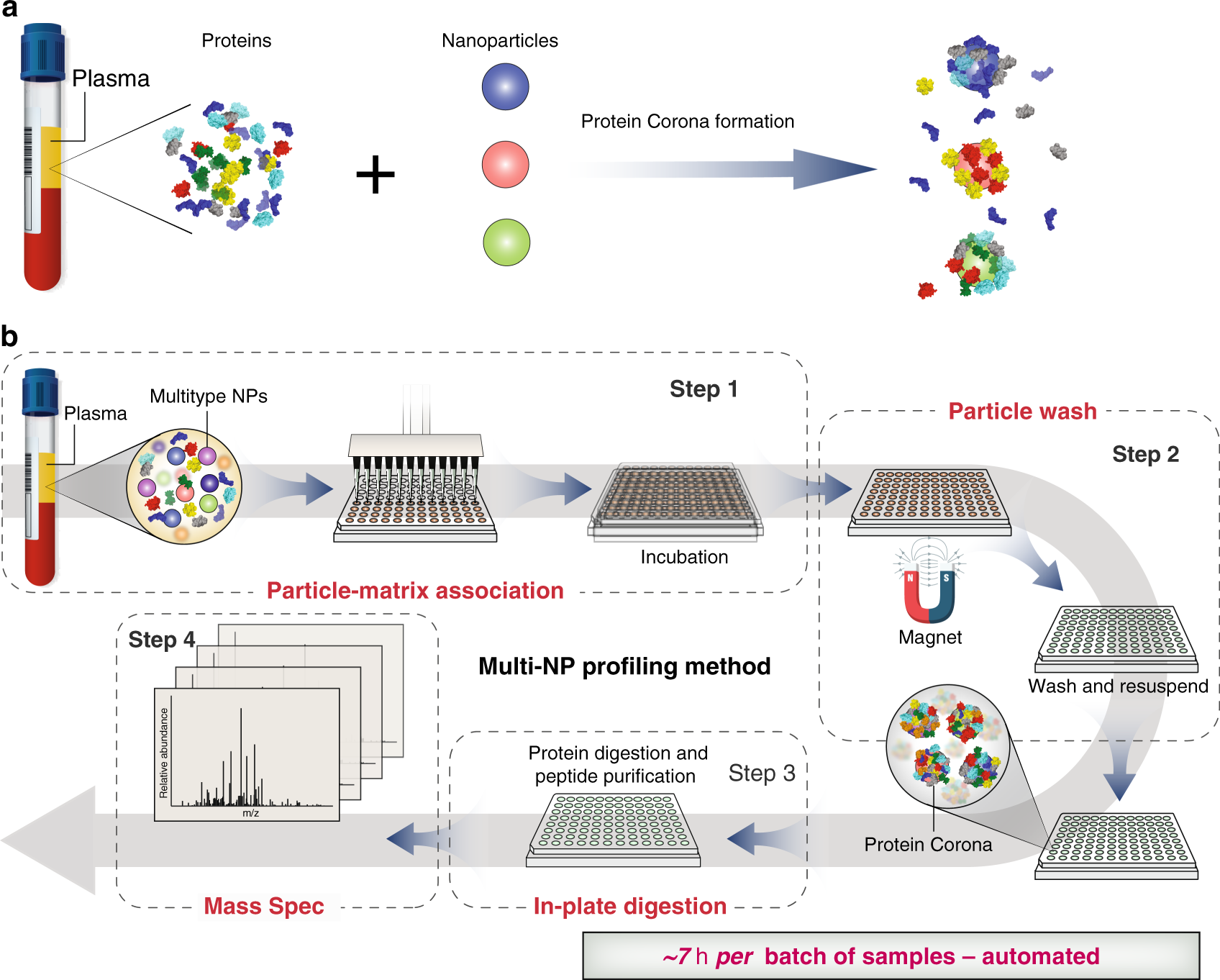
Rapid, deep and precise profiling of the plasma proteome with multi-nanoparticle protein corona | Nature Communications
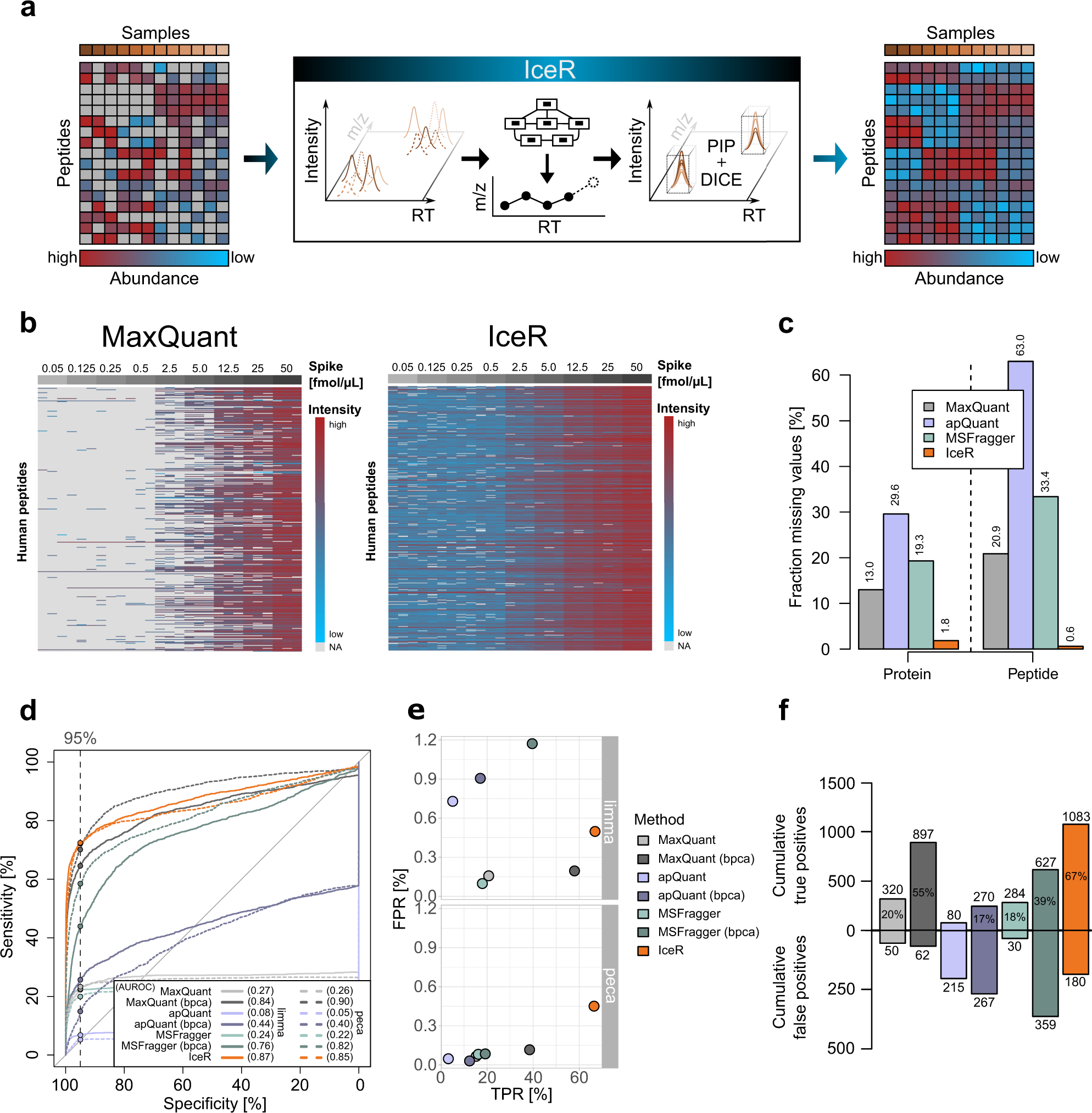
IceR improves proteome coverage and data completeness in global and single-cell proteomics | Nature Communications
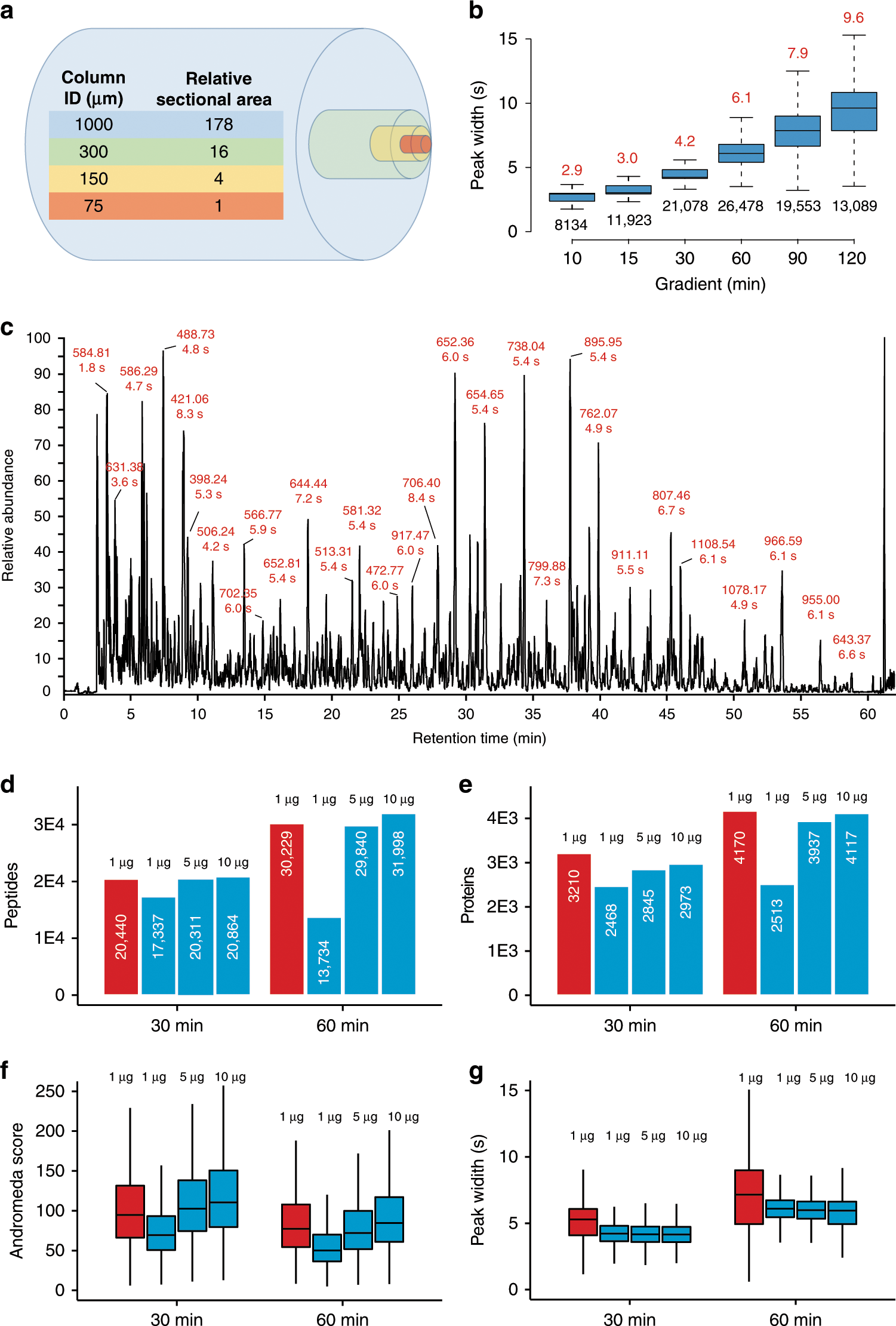
Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC–MS/MS | Nature Communications
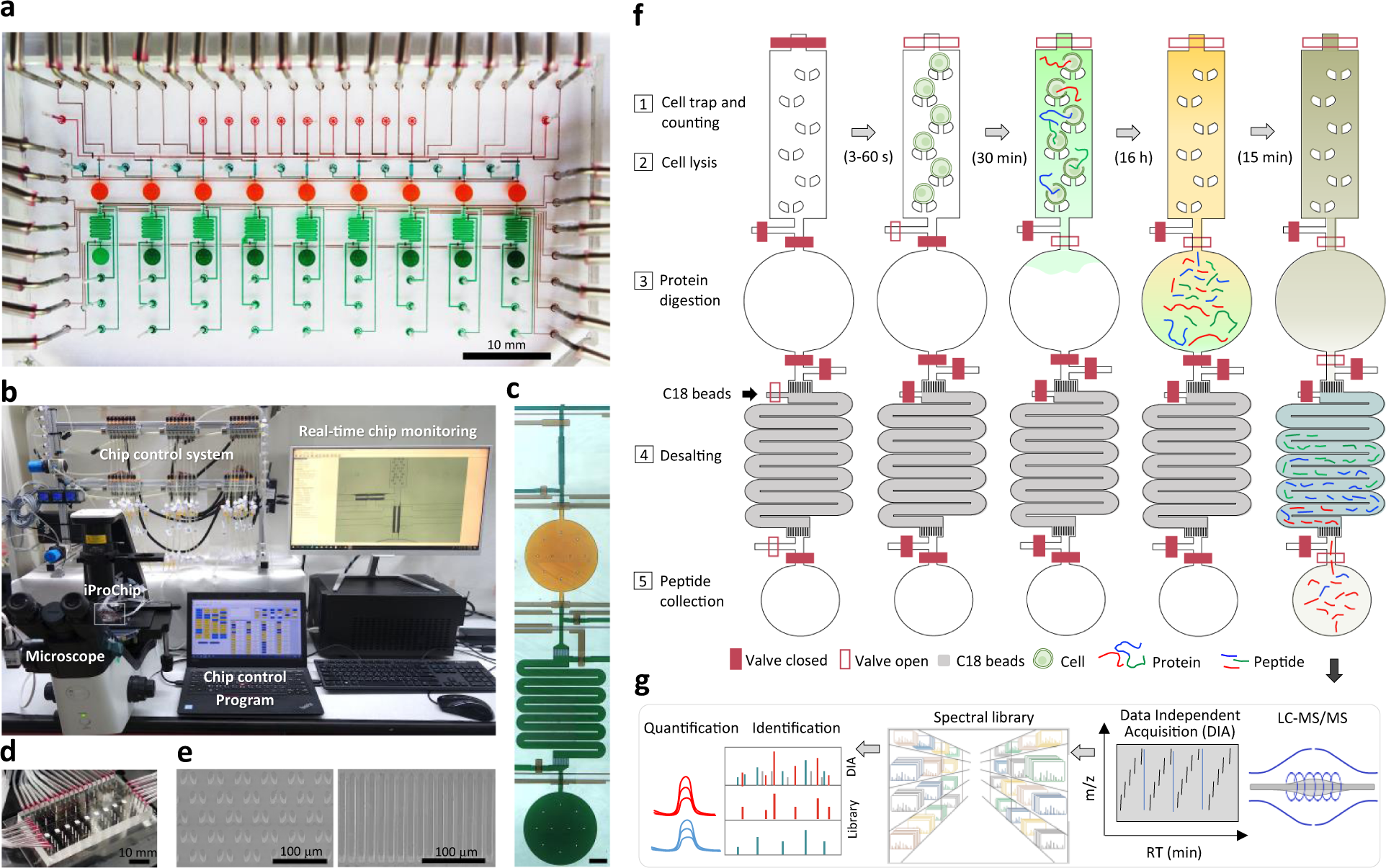
Streamlined single-cell proteomics by an integrated microfluidic chip and data-independent acquisition mass spectrometry | Nature Communications
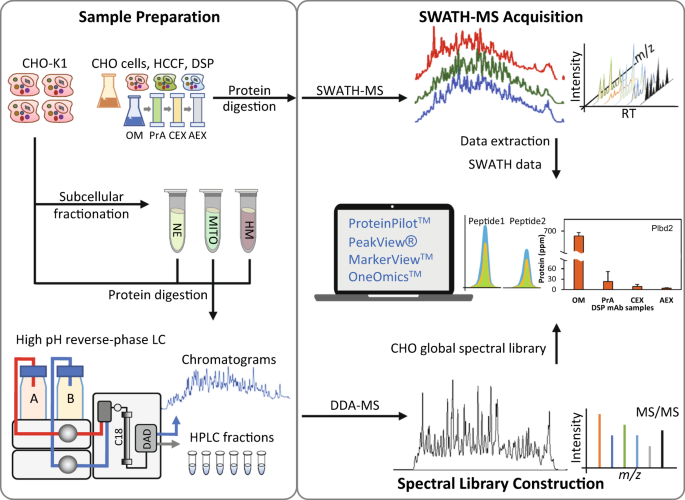
A comprehensive CHO SWATH-MS spectral library for robust quantitative profiling of 10,000 proteins | Scientific Data
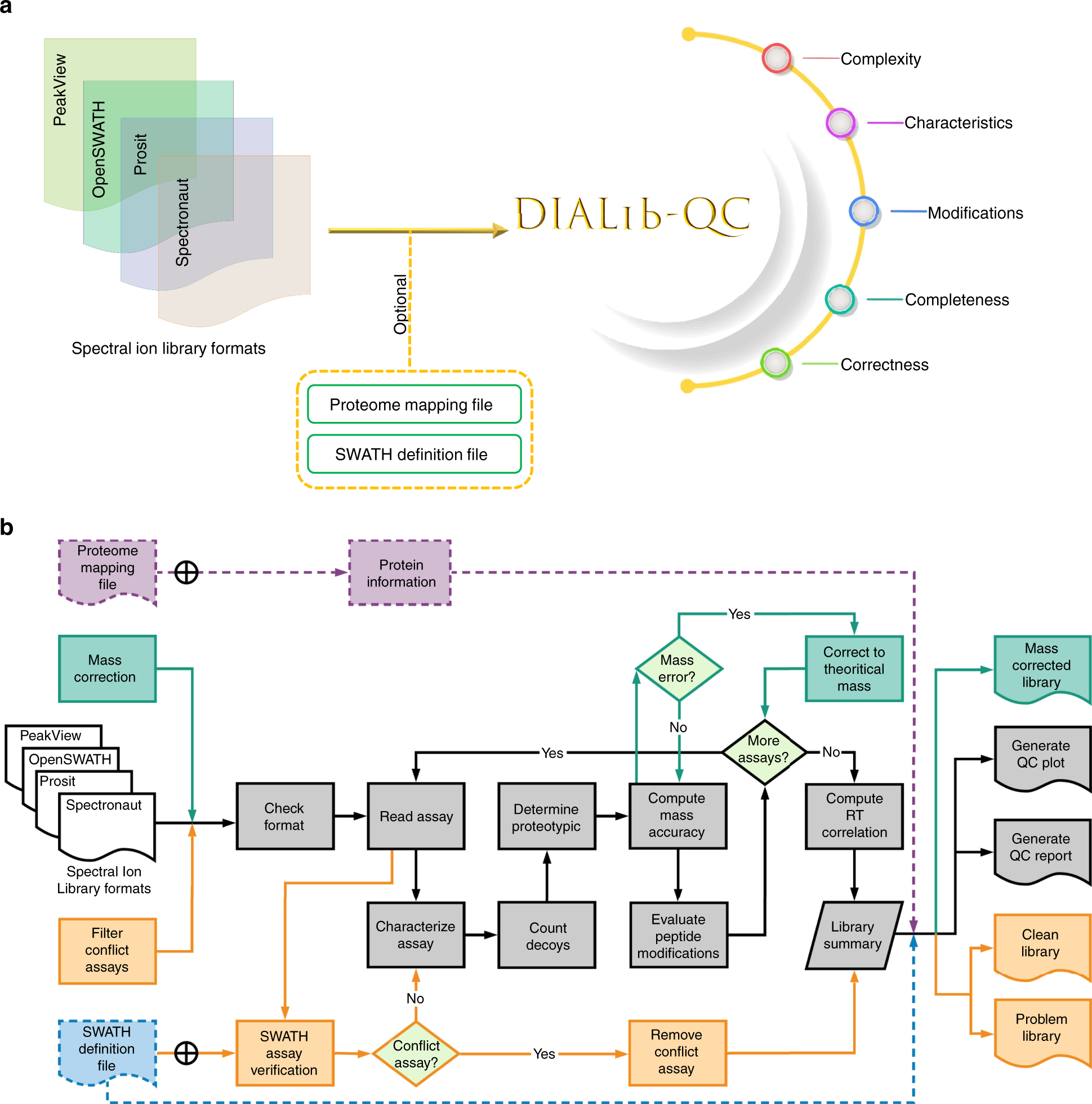
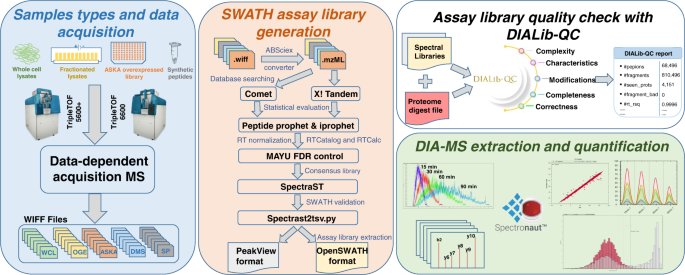
DIALib-QC an assessment tool for spectral libraries in data-independent acquisition proteomics | Nature Communications

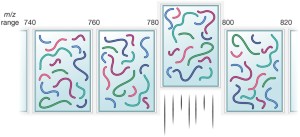
Technical comparison of DDA and different types of DIA in a biological... | Download Scientific Diagram

Deep Proteomics Using Two Dimensional Data Independent Acquisition Mass Spectrometry | Analytical Chemistry
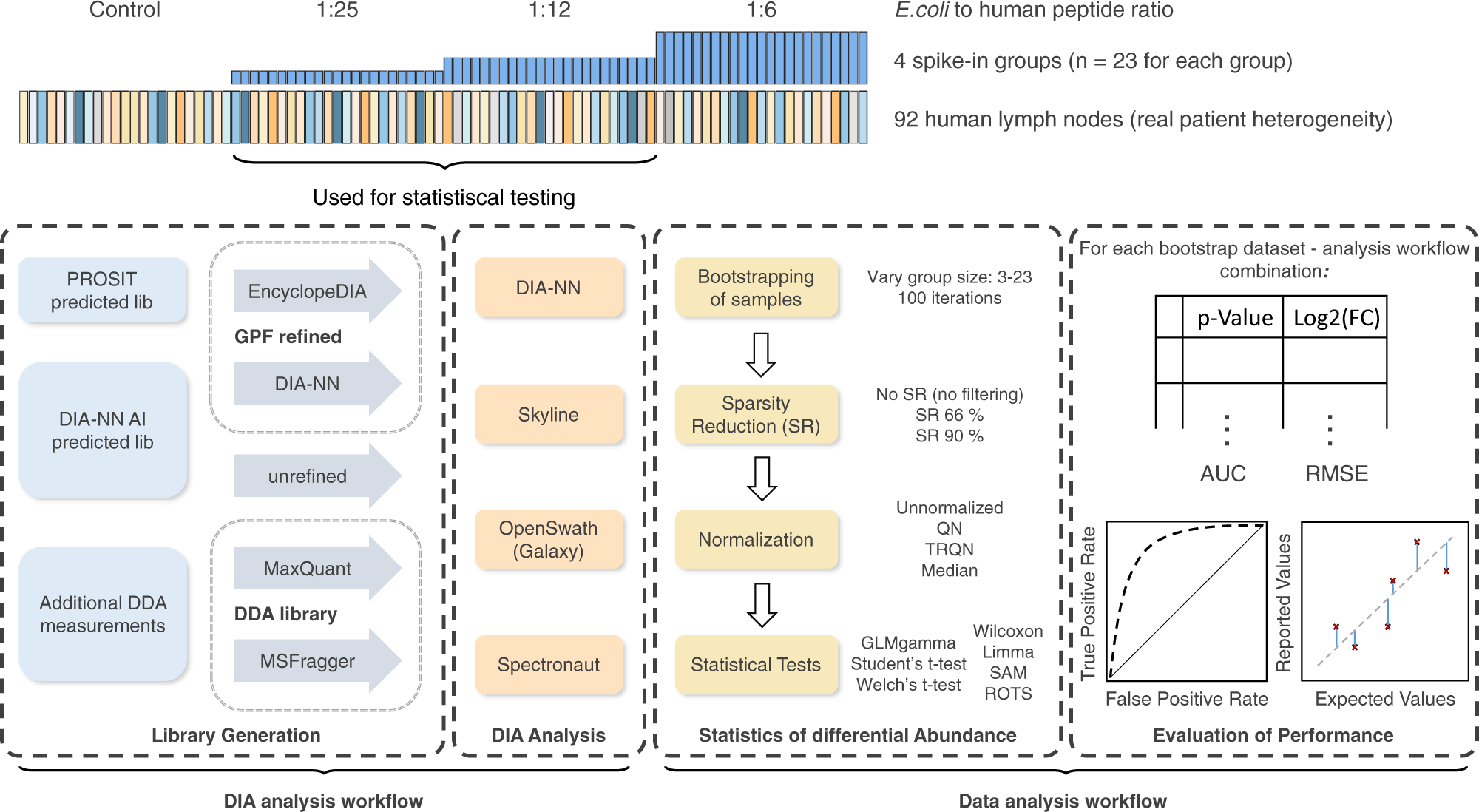
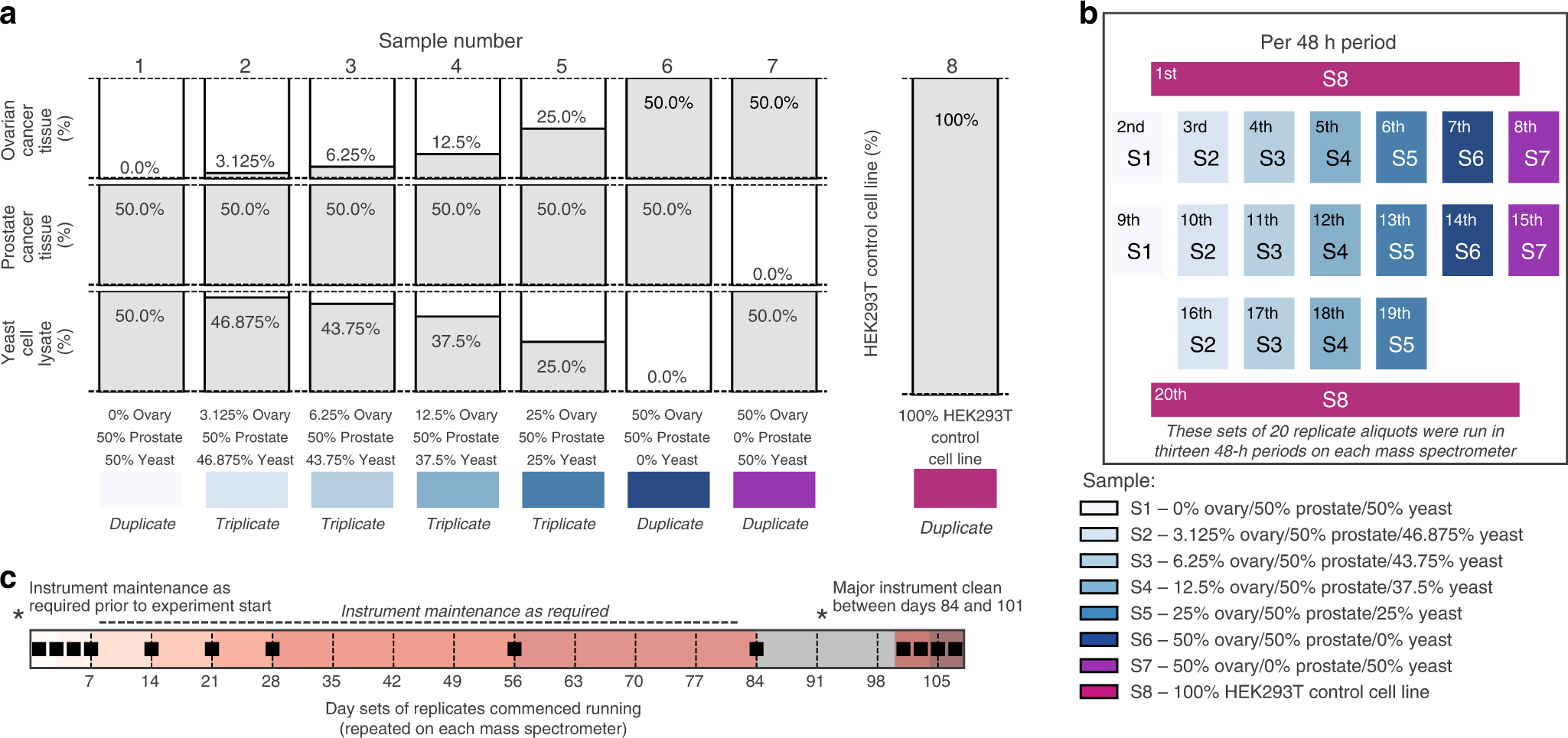
Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity | Nature Communications
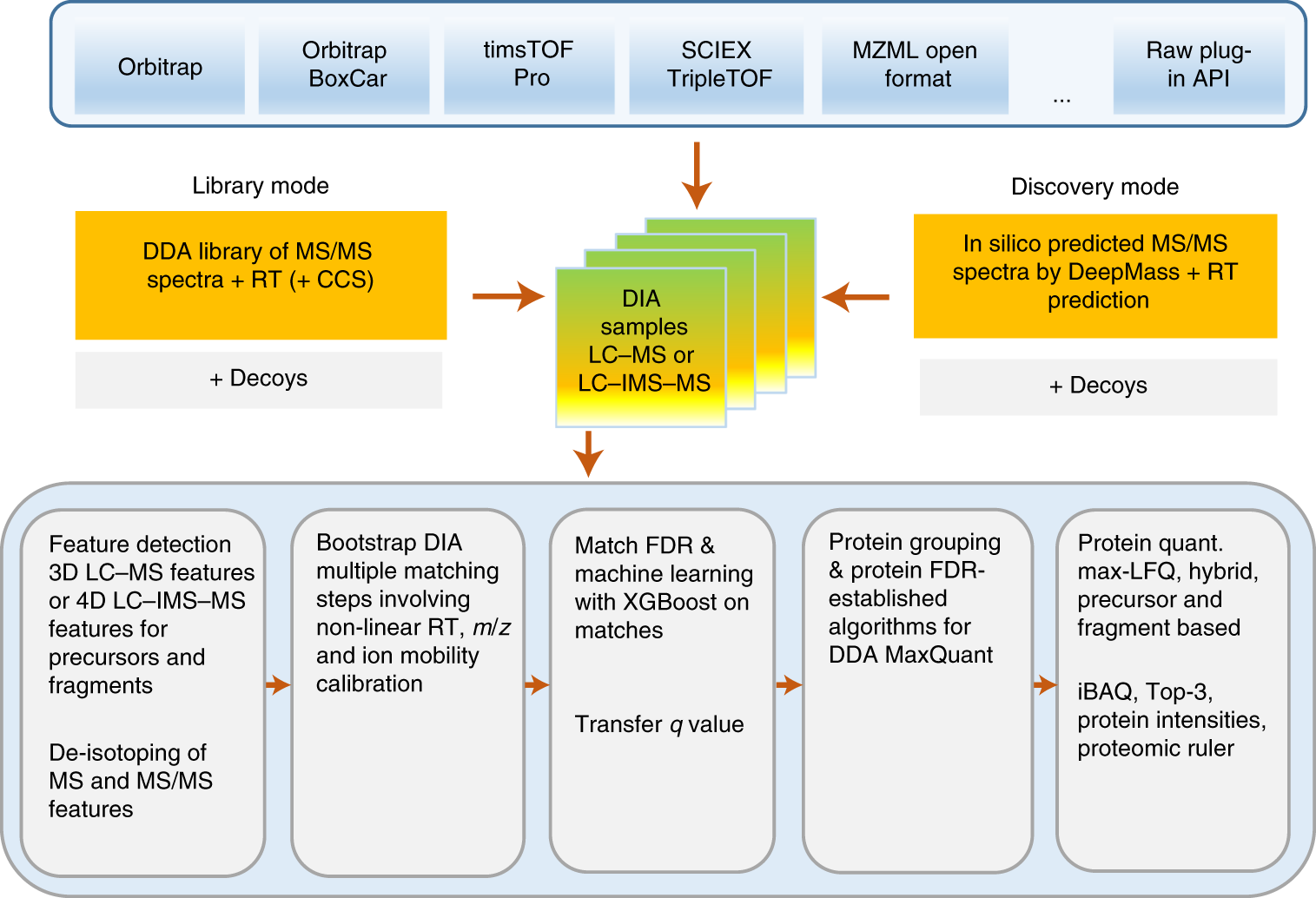
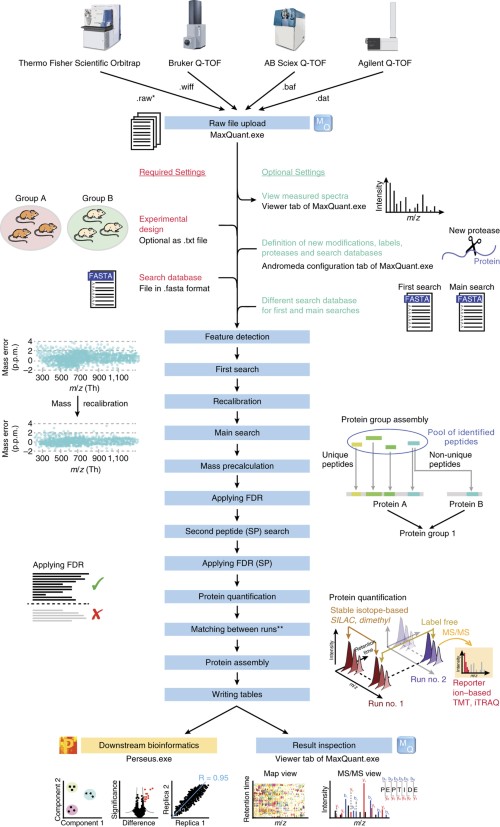
MaxDIA enables library-based and library-free data-independent acquisition proteomics | Nature Biotechnology
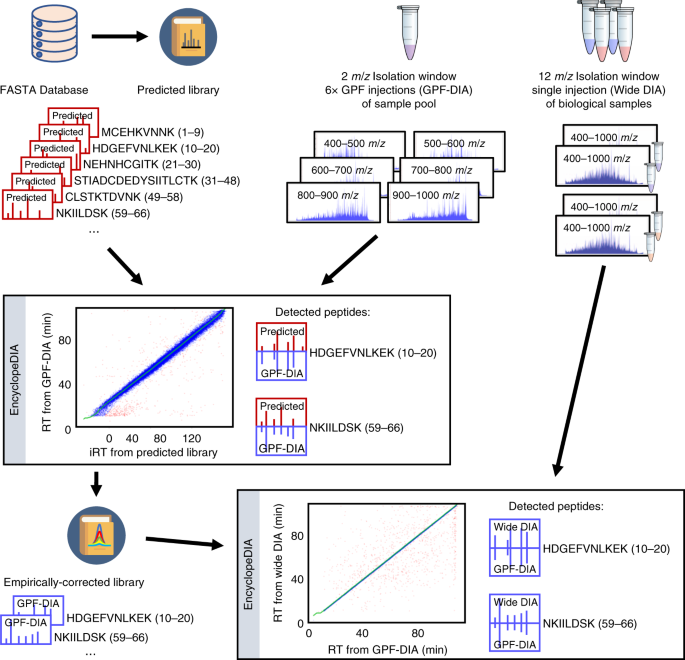
Generating high quality libraries for DIA MS with empirically corrected peptide predictions | Nature Communications
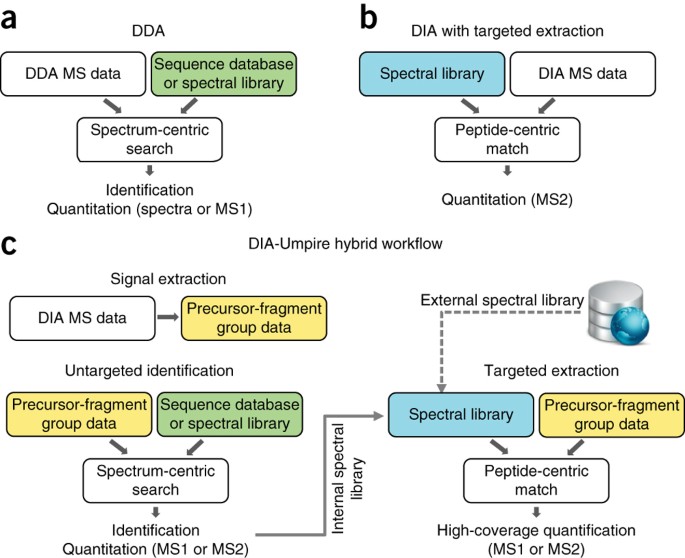
DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics | Nature Methods
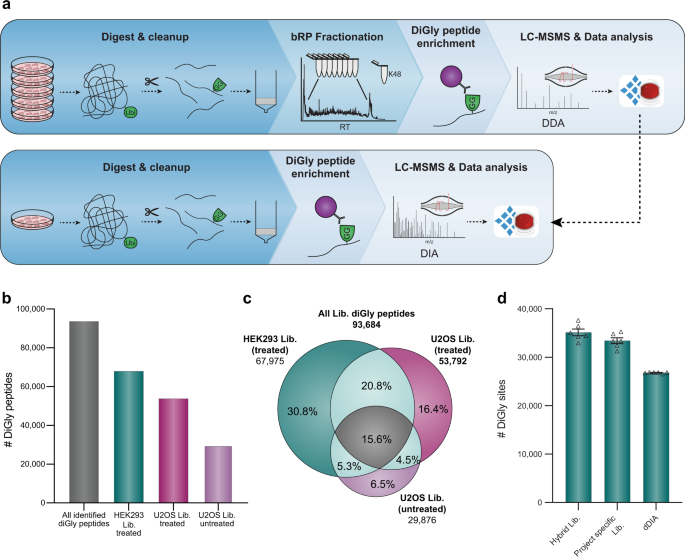
Data-independent acquisition method for ubiquitinome analysis reveals regulation of circadian biology | Nature Communications
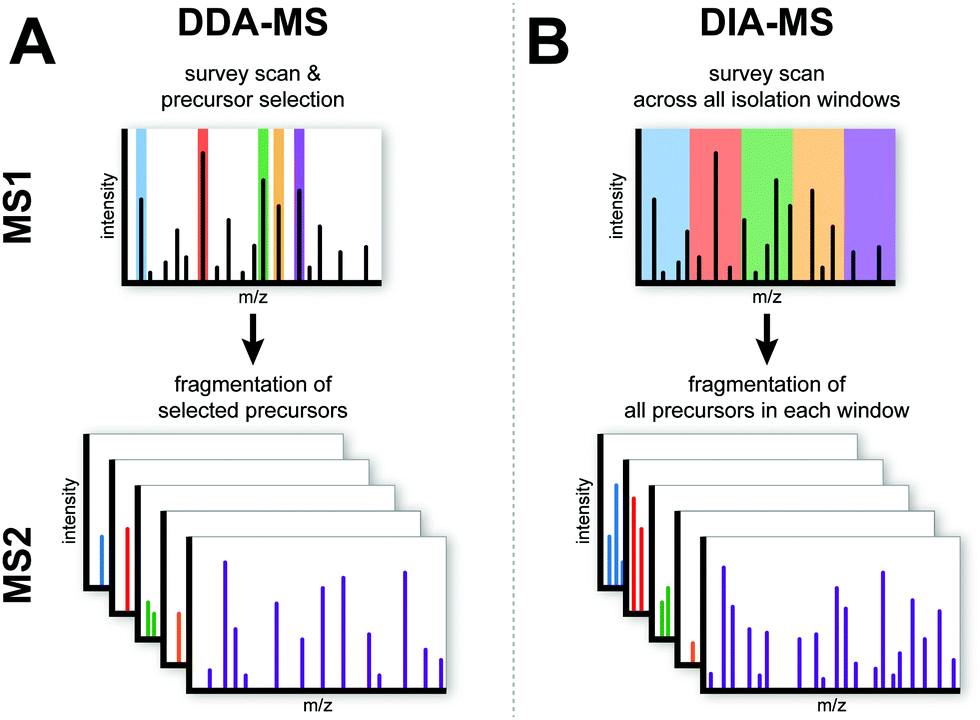
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H
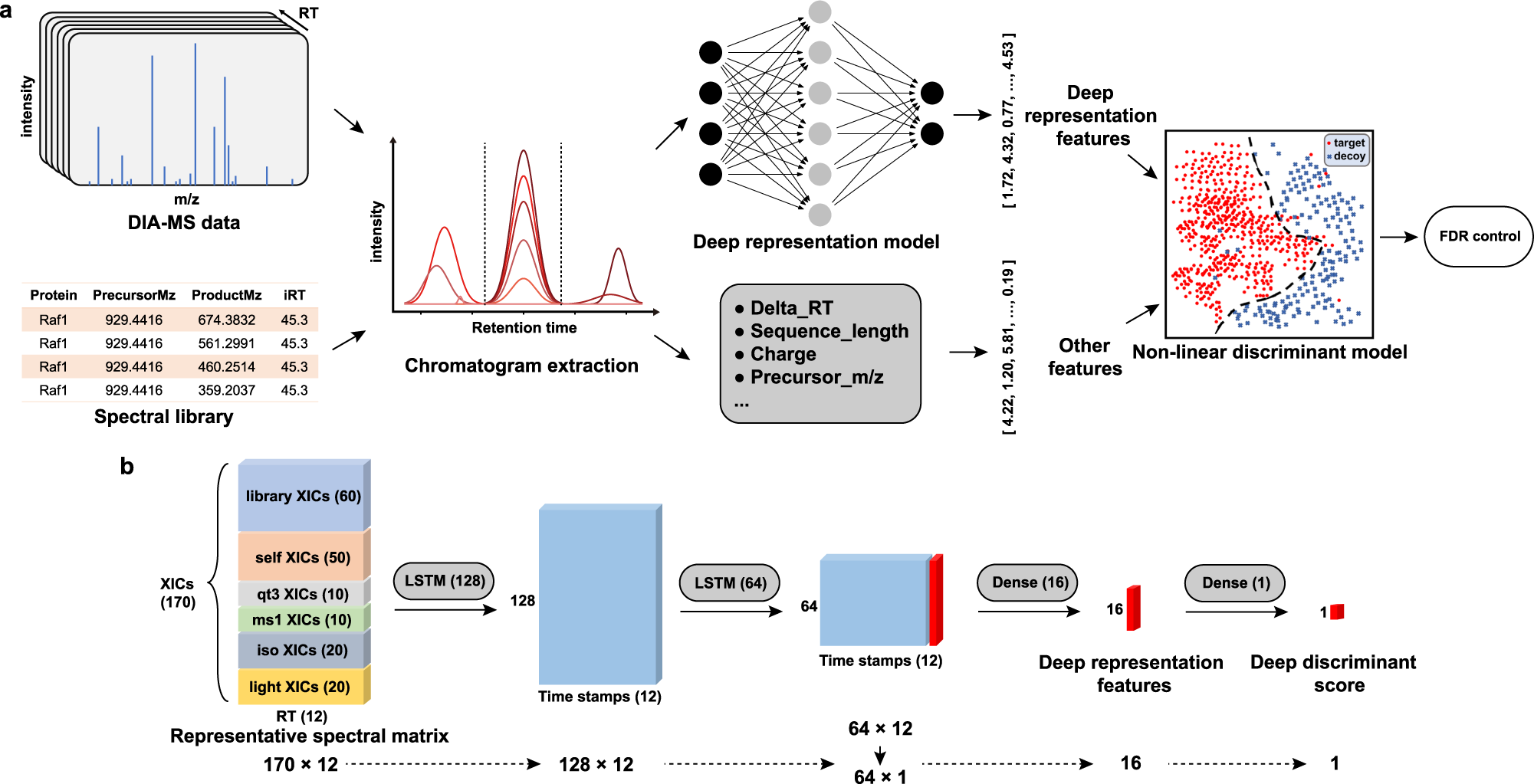
Deep representation features from DreamDIAXMBD improve the analysis of data-independent acquisition proteomics | Communications Biology

Hybrid Spectral Library Combining DIA-MS Data and a Targeted Virtual Library Substantially Deepens the Proteome Coverage - ScienceDirect
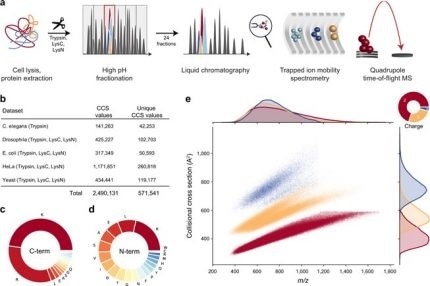
New Nature Communications publication by Mann & Theis Groups harnesses the benefits of large-scale peptide collisional cross section (CCS) measurements and deep learning for 4D-proteomics